Jason C. Hurlbert
Assistant Professor of Chemistry
Office: Sims 301B
Office hours: MWF 11-12 and by appointment
Phone: (803) 323-4928
email: hurlbertj@winthrop.edu
Research Interests
Our group is interested in studying the structure/function relationships of proteins. We use a variety of techniques to accomplish our goals but our research spans the fields of molecular biology, biochemistry and biophysics. Typically, we clone and express proteins in bacterial systems, purify the recombinant proteins, characterize them, idenitify conditions suitable for crystallization and then solve the structure of the crystallized protein. The variety of techniques used provides plenty of opportunities for undergraduates to learn techniques that they will need in whatever postbaccalaureate path they choose.
At present, our efforts are focused on two specific projects.
Structure and Function Studies of Human Sphingosine Kinases 1 & 2
Lipid signaling molecules such as ceramide, sphingosine, and sphingosine-1-phosphate (S1P) have recently been shown to be increasingly important in determining cellular fate. The sphingosine kinase family of proteins helps regulate the balance of these molecules in the cell1 and one of the two family members, sphingosine kinase 1 (SK1) is overexpressed in various types of cancers. Increased expression of SK1 has been associated with tumor angiogenesis and resistance to radiation and chemotherapy.
At present, we have developed a homology model of sphingosine kinase 1 (See images below) and our collaborator within the department, Christian Grattan, has used the model to aid in the design of inhibitors of the protein. However, the sequence differences between the bacterial lipid kinases whose structures are known and were used to build the model and the human sphingosine kinases are significant. A crystallographic structure is essential toward developing compounds with maximum therapeutic effectiveness.
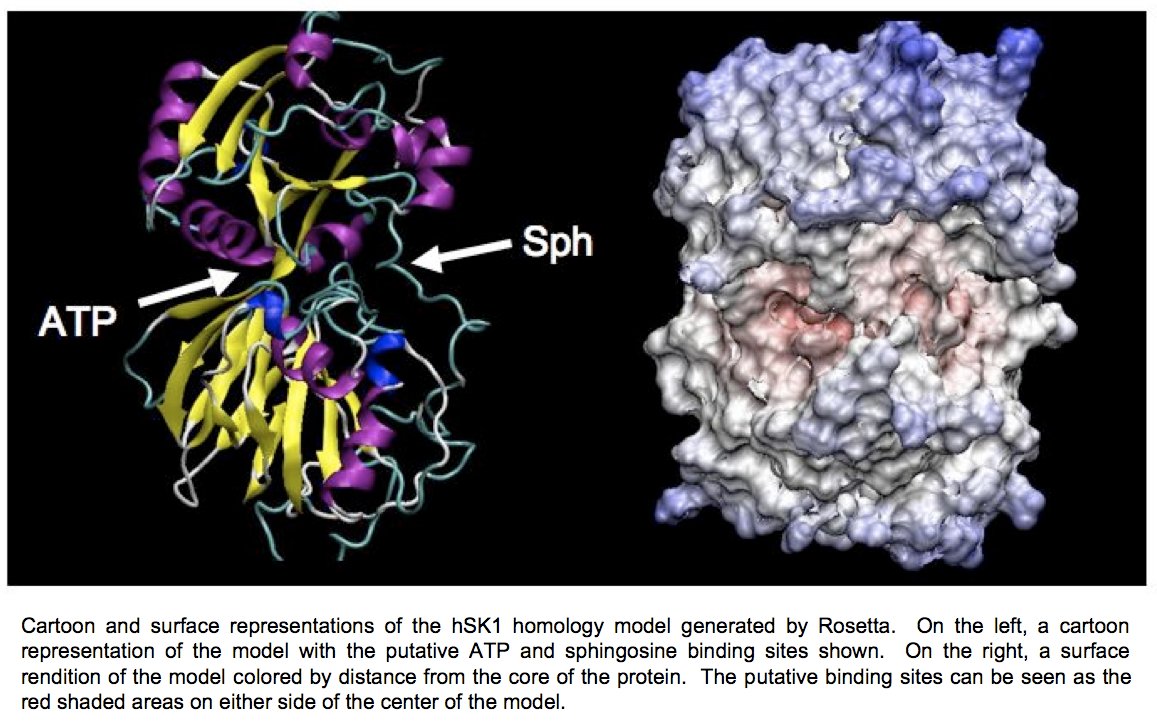
Determination of the three-dimensional structure of the proteins will allow for the crystallographic analysis of sphingosine kinases 1 and 2 in complex with substrates and biochemically verified inhibitors. Determination of the structure of this enzyme in complex with sphingosine, ADP, and previously characterized inhibitors will allow for a better understanding of the reaction mechanism of this novel family of enzymes.
Crystallographic Analysis of Xylanase C from Bacillus subtilis
Biodegradation of lignocellulosic biomass for the purpose of bioconversion to fuel ethanol is receiving renewed attention spurred by the high cost of petroleum and the desire for energy independence. Emphasis has been directed toward optimizing physical and chemical disruption and improvement of the enzymatic pretreatment of this disrupted biomass to maximize the release of fermentable sugars. New hydrolytic enzyme activities that promise to increase the efficiency of this process are of great interest. Biochemical characterization of xylanases of glycosyl hydrolase family five (GH 5) have revealed a novel mode of action that may contribute to these efforts.
Two GH 5 xylanases have been purified and studied for their biochemical properties. XynA from the Gram-negative bacterium Erwinia chrysanthemi and XynC from the Gram-positive bacterium Bacillus subtilis share 40% amino acid identity. Of primary interest in both studies was the specificity and mode of action on the complex substrate glucuronoxylan and the characterization of the xylooligomeric products generated. Both XynA and XynC are glucuronoxylan specific and cleave the β-1,4 linked xylan main chain in an endo manner directed by the position of an α-1,2 linked glucuronate moiety. Results from several biochemical techniques all support the conclusion that cleavage occurs at the second β-1,4 linkage toward the reducing terminus from a glucuronate substituted xylose. This mode of action results in singly substituted glucuronoxylose oligosaccharide products with the glucuronic acid substitution penultimate to the reducing terminus (Figure 1). The novel specificity sets these enzymes apart from the more common endo-acting xylanases that cleave the xylan chain in assessable regions. This unique mode of action identifies the first endoxylanase family that degrades polymeric xylan in a directed rather than a random manner.
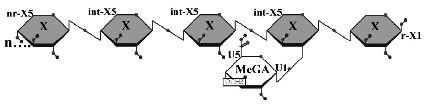
Figure 1: Limit product generated by the hydrolysis of glucuronoxylan by B. subtilis XynC. X, xylose; MeGA, 4-O-methylglucuronic acid; n, some number of beta-1,4-linked xylose residues.
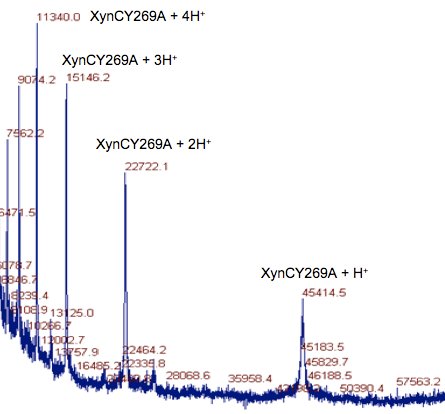
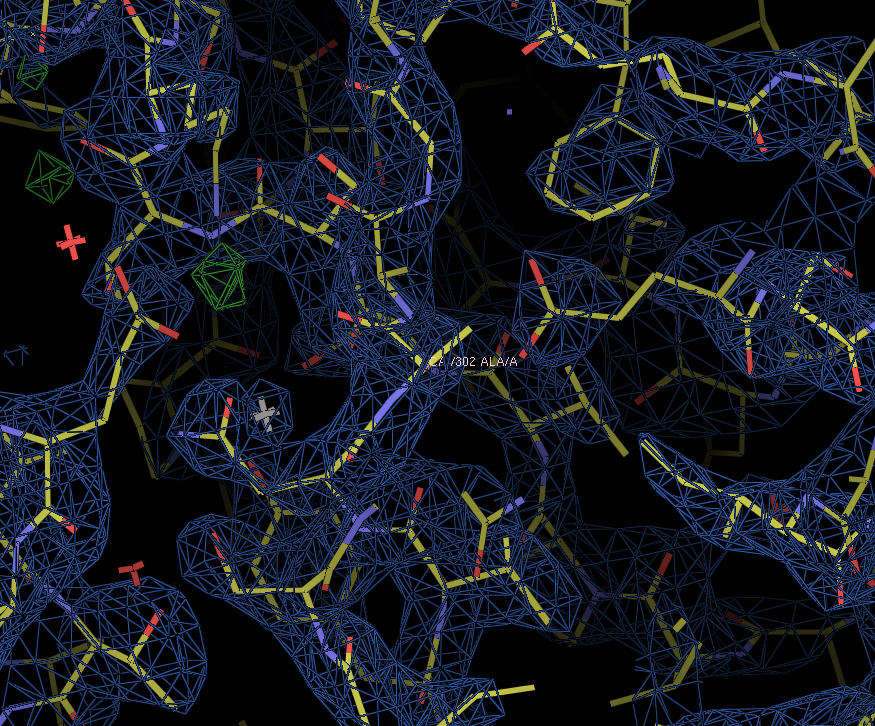
XynC is the second GH 5 xylanase to be structurally studied. Recently, the high resolution crystallographic structure of XynC has been determined resulting from a collaborative effort involving the principal investigator, a student researcher, David Godwin, from Winthrop University and Dr. Franz St. John at the University of Maryland. Samples of recombinant protein were produced, purified and crystallized at Winthrop and then sent to the University of Maryland for crystallographic data collection. Diffraction data were collected on protein crystals using an x-ray source at the University of Maryland as well a portion of their time on a beamline at the Stanford Synchrotron Radiation Laboratory. These data were then analyzed by the principal investigator and the structure of the protein was solved to 1.8Å with molecular replacement techniques (Figure 2). The resulting structure has allowed for subsequent rapid determination of the structure of the enzyme in complex with a variety of oligoxylosides, oligomeric glucuronoxylosides and glucuronic acid moieties. Unfortunately, the electron densities of the oligosaccharides were unable to be resolved; most likely due to molecular motion and/or incomplete occupancy of the ligands in the active site. The structure of the enzyme in complex with glucuronic acid was able to be determined to 2.8Å with a clear density present for the pyranose ring. This structure provides direct evidence for the observed specificity of the enzyme and serves as the basis for the rationale of the current proposal.
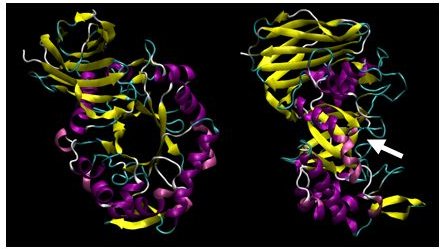
Figure 2: X-ray crystallographic structure of B. subtilis XynC at 1.8Å resolution. The model is colored such that alpha helices are purple and beta strands are colored yellow. The model on the right is a 90 degree rotation of the figure on the left. The active site cleft is indicated by the arrow.
Using the structure of XynC complexed with glucuronic acid, we are proposing to identify the role of several amino acids surrounding the active site in the demonstrated selectivity of the enzyme. We will accomplish this goal by a combination of computational, molecular biological and biophysical techniques including: in silico docking studies, site-directed mutagenesis, recombinant DNA expression, chromatographic separation of proteins, enzymological analysis, protein crystallization and x-ray diffraction analysis. Ultimately, we aim to clearly identify the role of tyrosines 231, 269 and 274, serine 235 and arginine 272 in the binding of glurcuronic acid substituted xylan substrates. This information will then be used to further explain the function of the glycosyl hydrolase family 5 xylanases.
Publications
St John, F.J., Hurlbert, J.C., Rice, J.D., Preston, J.F. and Pozharski, E., 2011. Ligand Bound Structures of a Glycosyl Hydrolase Family 30 Glucuronoxylan Xylanohydrolase. J. Mol. Biol. 407: 92109.
St John, F.J., Godwin, D.K., Preston J.F., Pozharski E. and Hurlbert, J.C., 2009. Purification, crystallization and crystallographic analysis of Bacillus subtilis xylanase C Acta Cryst. F65: 499–503
Zhang, Y.-M., Hurlbert, J. C., White, S. W., Rock, C. O., 2006. Roles of the active site water, histidine 303, and phenylalanine 396 in the catalytic mechanism of the elongation condensing enzyme of Streptococcus pneumoniae. Journal of Biological Chemistry 281: 17390-9.
Preston, J.F., J.C. Hurlbert, J.C. Rice, A. Ragunathan, and F.J. St. John, 2003. "Chapter 12: Microbial Strategies for the Depolymerization of Glucuronoxylan: Leads to the Biotechnological Applications of Endoxylanases." ACS Symposium Series 855, Applications of Enzymes to Lignocellulosics. Edited by Shawn D. Mansfield and John Saddler.
Hurlbert, J.C. and T. Izard, 2002. Crystallization of HLA-DR4 Complexed with a Collagen II Peptide. Acta Crystallographica D. 58: 1749-1751.
Hurlbert, J.C. and J.F. Preston, 2002. Differences in the solution structures of the parallel β-helical pectate lyases as determined by limited proteolysis. Biochimica et Biophysica Acta - Proteins and Proteomics 1599: 9-20.
Hurlbert, J.C. and J.F. Preston, 2001. Functional characterization of a novel xylanase from a corn strain of Erwinia chrysanthemi. Journal of Bacteriology 183: 2093-2100.
Hurlbert, J.C. and J.F. Preston, 2000. Functional implications of the β-helical protein fold: Differences in chemical and thermal stabilities of Erwinia chrysanthemi EC16 pectate lyases B, C and E. Archives of Biochemistry and Biophysics 381: 264-272.
Presentations
Bush, B., Hurlbert, J., Sumter, T.F. Regulation of High Mobility Group A1 expression by the Adenomatous polyposis coli (Apc) tumor suppressor gene. Poster presented at the 2009 Meeting of the American Society of Biochemistry and Molecular Biology (ASBMB), April 19, 2009, New Orleans, LA.
White, E., Grattan, T.C. and J. Hurlbert. Design of SKI-I Derivatives through in Silico Docking for Improved Oral Bioavailability. Poster presented at the 60th Southeastern Regional Meeting (SERMACS), November 12, 2008, Nashville, TN.
Parker, C., Ghadiyali, Z., Kissinger, S., MckInney. K and J. Hurlbert, 2007. Expression and Purification of Human Adiponectin Membrane Receptors, AdipoR1 and AdipoR2. Poster presented at the Experimental Biology 2007 meeting, Washington, D.C.
Glazener, R., Baker, I., Hurlbert, J. and C. Parker, 2007. Kinetic characterization of Arylamine N-acetyltransferase using the substrates sulfamethazine and 5-aminosalicylic acid. Poster presented at the 233rd American Chemical Society National Meeting, Chicago, IL.
Glazener, R., Baker, I., Hurlbert, J. and C. Parker, 2007. Kinetic characterization of arylamine-N-acetyltransferase 2. Poster presented at the 21st National Conference on Undergraduate Research, San Rafael, CA.
Kissinger, S., Walker, L., Hurlbert, J. and C. Parker, 2007. Expression of Human Adiponectin Membrane Receptor 1 in Sf9 cells. Poster presented at the 21st National Conference on Undergraduate Research, San Rafael, CA.
Hurlbert, J.C. and Tina Izard, 2002. Crystallization of HLA-DR4 complexed with a collagen II peptide. Poster presented at the 9th International Convention on the Crystallization of Biological Macromolecules, Jena, Germany.
Preston, J.F., Rice, J.D. and Jason C. Hurlbert, 2002. Microbial strategies for the depolymerization of methylglucuronoxylan. Poster presented at the 223rd American Chemical Society National Meeting, Orlando, Florida.
Hurlbert, J.C. and J.F. Preston, 1999. Thermal and chemical denaturation of pectate lyases B, C and E secreted by Erwinia chrysanthemi EC16. The FASEB Journal 13: p. A1571.
Hurlbert, J.C. and J.F. Preston, 1998. Differences in solution structure of pectate lyases C and E determined by limited proteolysis. The FASEB Journal 12: p. A1450.
Hurlbert, J.C. and J.F. Preston, 1996. Comparative properties of pectate lyases B and C secreted by Erwinia chrysanthemi EC16. Abstracts of the Annual Meeting of the American Society for Microbiology. Abstract K175.
Preston, J.F., Y.T. Chen, J.D. Rice and J.C. Hurlbert, 1994. Diversity of alginolytic activities produced by Vibrio sp. Associated with the brown algae, Sargassum fluitans. Abstract for the First International Conference on Polysaccharide, Trondheim, Norway. p. A-10
|

